install.packages("LIBRARY")1 Getting Started with R
1.1 Your R System
In this course, we work with the combination of R + RStudio.
- R is the calculation engine (language and environment) that performs the computations. You can download the latest R version here.
- RStudio is the editor (IDE) that helps you sending inputs to R and collect outputs. You can download the latest Rstudio version here.
Make sure you have a recent version of R + RStudio installed on your computer.
If you have never used RStudio, here is a good video introducing the basic system and how R and RStudio interact, and here you have simple and good explanations for each Rstudio panel and the many resources.
1.2 Libraries that you will need
The R engine comes with a number of base functions, but one of the great things about R is that you can extend these base functions by libraries that can be programmed by anyone. In principle, you can install libraries from any website or file. In practice, however, most commonly used libraries are distributed via two major repositories. For statistical methods, this is CRAN, and for bioinformatics, this is [bioconductor](https://www.bioconductor.org/).
To install a package from a library, use the command:
Exchange “LIBRARY” with the name of the library you want to install. The default is to search the package in CRAN, but you can specify other repositories or file locations in the function. For Windows / Mac, R should work out of the box. For other UNIX based systems, may also need to install
build-essential
gfortran
libmagick++-dev
r-base-dev
cmakeIf you are new to installing packages on Debian / Ubuntu, etc., type the following:
sudo apt update && sudo apt install -y --install-recommends build-essential gfortran libmagick++-dev r-base-dev cmakeIn this book, we will often use data sets from the EcoData package, which is not on CRAN, but on a [GitHub page](https://github.com/TheoreticalEcology/ecodata). To install the package, if you don’t have the `devtools` package installed already, first install devtools from CRAN by running:
install.packages("devtools")Then, install the EcoData package via:
devtools::install_github(repo = "TheoreticalEcology/EcoData",
dependencies = T, build_vignettes = T)For your convenience, the EcoData installation also forces the installation of most of the packages needed in this book, so this may take a while. If you want to load only the EcoData package, or if you encounter problems during the install, set dependencies = F, build_vignettes = F.
If the installation didn’t work, download the package file manually from:
https://github.com/TheoreticalEcology/ecodata/releases/download/v0.2.1/EcoData_0.2.1.tar.gz
Store the file on your computer in the same folder where you created your R project. Then run the following code:
install.packages("EcoData_0.2.1.tar.gz",
repos = NULL, type = "source")
library(EcoData)1.3 Setting up the working environment in RStudio
Your first task is to open RStudio and create a new project for the course.
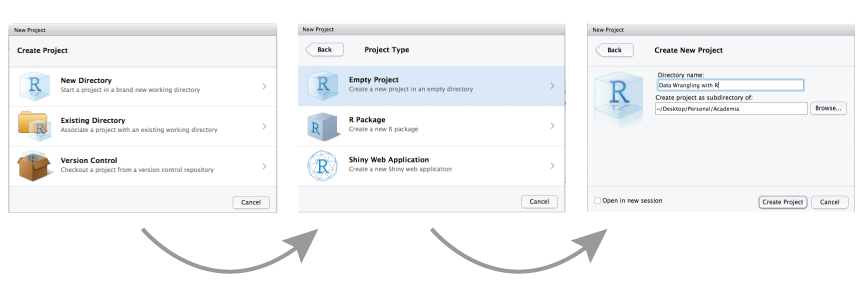
- Click the ‘File’ button in the menu, then ‘New Project’ (or the second icon in the bar below the menu “Create a project”).
- Click “New Directory”.
- Click “New Project”.
- Type in the name of the directory to store your project, e.g. “IntroStatsR”.
- “Browse” to the folder on your computer where you want to have your project created.
- Click the “Create Project” button.
For all exercises during this week, use this project! You can open it via the file system as follows (please try this out now):
- (Exit RStudio).
- Navigate to the directory where you created your project.
- Double click on the “IntroStatsR.Rproj” file in that directory.
You should now be back to RStudio in your project.
In the directory of the R project, generate a folder “scripts” and a folder “data”. You can do this either in the file directory or in RStudio. For the latter:
- Go to the “Files” panel in R Studio (bottom right panel).
- Click the icon “New Folder” in the upper left corner.
- Enter the folder name.
- The new folder is now visible in your project directory.
The idea is that you will create an R script for each exercise and save all these files in the scripts folder. You can do this as follows:
- Click the “File” button in the menu, then “New File” and “R Script” (or the first icon in the bar below the menu and then “R Script” in the dropdown menu).
- Click the “File” button in the menu, then “Save” (or the “Save” icon in the menu).
- Navigate to your scripts folder.
- Enter the file name, e.g. “Exercise_01.R”.
- Save the file.